What is Epigenetics technical articles are geared towards epigenetic research techniques, news, and trends in the field of epigenetics, written by scientists from universities and institutions including UCLA, Hofstra, NIH, Johns Hopkins, and more.
Explore in detail new epigenetic research techniques and tips for topics like next-generation sequencing (NGS) in epigenetics, m6A RNA methylation, CRISPR/Cas9 system epigenetic editing, chromatin immunoprecipitation (ChIP) protocol optimization, and single-cell epigenomics methods.
Want to share your research? Submit your own epigenetics article to be featured.
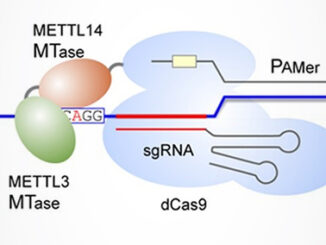
“The fifth RNA base” N6-methyladenosine, or m6A, is the most common and abundant eukaryotic RNA modification, accounting for over 80% of all RNA methylation. It can be found mainly in mRNA, but is also observed in non-coding species like tRNA, rRNA, and miRNA. Through interactions with various binding proteins called “readers”, m6A affects virtually every facet of ribonucleic acid biology: structure, splicing, localization, translation, stability, and turnover [1]. Aside from this central role in RNA metabolism, m6A is a factor [more…]
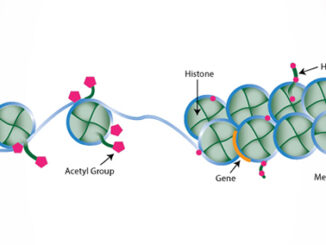
This article was first published by EpigenTek. In the early 1960s, Vincent G. Allfrey and his colleagues at the Rockefeller Institute (today’s Rockefeller University) demonstrated by way of C14 labeling the incorporation of methyl and acetyl groups into histones.1 The evidence suggested that these chemical additions succeeded protein synthesis, although the means by which they were incorporated as well as the extent of their biological significance were not fully understood at the time. Allfrey’s pioneering work effectively laid the foundation for histone [more…]
Decades of antibiotic and antifungal use have dramatically impacted the global microbiome, with ramifications for agriculture and human health. Worries are mounting but no systematic plans of action are forthcoming. Though that is not to say there are no slivers of hope. While they are still largely confined to immunocompromised patients, fungal infections are a looming public health crisis. Invasive candidiasis is increasingly caused by multidrug-resistant non-albicans yeasts and nearly 60% of C. parapsilosi sampled in South Africa are now [more…]
For centuries, scientists and philosophers have been fascinated with learning about the brain. However, it hasn’t been until the last few decades that we’ve really been able to examine this complex organ in action. One advanced technology that has revolutionized our understanding of the human brain is magnetic resonance imaging (MRI). While this method has been extremely useful for measuring neural activity, it hasn’t been used to assess gene regulation – that is, until now. The University of Illinois (U [more…]
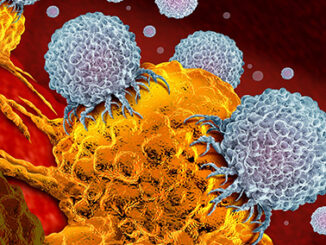
The body’s first line of defense against infection is the innate immune system. Although it is non-specific and less effective than the acquired immune system, it responds to invading pathogens much faster, holding off an immunological threat until a specific response can be mobilized. One integral part of the innate immune system is natural killer cells (NK). As their name suggests, these cells are capable of killing affected cells autonomously while also signaling an immune response as they secrete pro-inflammatory [more…]
A new technique has been developed that is expected to advance our knowledge of numerous underlying biological processes, including those implicated in complex diseases like cancer. Using machine learning —a form of artificial intelligence— scientists can predict gene regulation at the cellular level – a process that, before now, has been nearly impossible to do accurately. An important goal in epigenetic research is to identify regions in the genome that are vulnerable to molecular factors that can alter gene expression [more…]
Histones are the most eminent DNA-interacting proteins. As the primary protein constituent of chromatin, forming complexes with DNA to compact our large genome for efficient nuclear organization, histones support critical cellular processes such as transcription, DNA replication, and DNA repair through diverse post-translational modifications that regulate their interactions with DNA and other nuclear proteins. Consequently, anomalous changes in histone epigenetics can lead down the road to assorted pathological conditions, making histone modifications and their associated modifying enzymes suitable diagnostic, prognostic, [more…]
Cells are continually being generated and eliminated by our bodies in an ongoing cycle crucial for maintaining proper tissue structure and function. The death of cells during normal tissue maintenance results in the release of intracellular DNA material into the bloodstream, where it freely circulates. In fact, healthy individuals can have up to 100 nanograms of non-encapsulated DNA fragments floating around in every milliliter of their blood.1 Circulating cell-free DNA, or cfDNA for short, has proven quite useful in various [more…]
Antibodies are used extensively in research for procedures, such as western blotting (WB), enzyme-linked immunosorbent assays (ELISA), and immunoprecipitation (IP). The issue of accuracy and reproducibility surrounding the usage of commercially available antibodies for research continues to be a concern within the scientific community. The best way to compile information about an antibody is to review the provided manufacturer datasheet. In fact, some journals will not publish a study that includes an antibody with insufficient information and validation effort. Researchers [more…]
The most common epigenetic mark throughout the genome is 5-methylcytosine (5mC). Additionally, 5-hydroxymethylcytosine (5hmC) is also known to play a role in various processes, including embryonic stem cell development and tumorigenesis. There is also an enrichment of 5hmC in human neurons. Accurately quantifying and determining the genomic location of these two different marks is therefore crucial to our understanding of molecular biology and disease. Standard bisulfite sequencing (BS-seq) has been a reliable method for detecting un-methylated and methylated cytosine residues [more…]