
Neurogenesis is a highly coordinated process with sequential waves of tightly controlled changes in gene expression. N6-Methyladenosine (m6A), is believed to play a role in the epitranscriptomic mechanism that controls mammalian cortical neurogenesis.
M6A is the most abundant modification in mRNA and is found in most eukaryotic cells. It is installed by the methyltransferase complex, consisting of methyltransferase-like 3 (Mettl3) and methyltransferase-like 14 (Mettl14) as the SAM-binding sub-unit. The role of m6A modification in cancer has previously been investigated. Now, the Yoon et al. (2017) study demonstrates that m6A methylation plays a conserved regulatory role in cortical neurogenesis in both humans and mice.
In this study, Mettl14 conditional knockout (cKO) mice were used to examine m6A function in embryonic cortical neurogenesis. A Mettl14 deletion resulted in the cKO mice dying before postnatal day 25 (p25), suggesting that m6A is vital for nervous system development.
Immunohistology was then performed on the postnatal day 5 (p5) cKO mice in order to determine how exactly m6A affects the development of the nervous system. At p5, the cKO mice exhibited enlarged ventricles with an adjacent dense layer of cells that resembled the embryonic germinal zone that differed from their WT littermates.
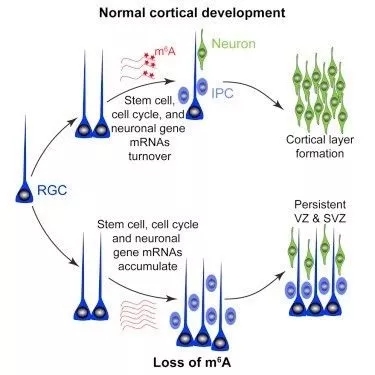
Pax6+ and Nestin+ cells presence was also observed in cKO mice but not WT. During normal mouse cortical development, Pax6+ radial glial cells (RGCs), are depleted. This difference in development suggests that the cKO mice maintain neurogenesis into postnatal stages. The team later confirmed this when they compared the number of EdU+Satb2+ neurons between cKO and WT mice. The decreased number of EdU+Satb2+ neurons suggests a deficit in producing late-born upper-layer neurons.
Given the differences in development timing between cKO and WT mice, the interkinetic nuclear migration in phase with cell-cycle progression was observed. When they compared the cKO mice to the WT mice at the 6 hr mark, the RGCs in the cKO mice were positioned further away from the ventricular surface than in WT mice. This suggests that the interkinetic nuclear migration of the cKO mice is delayed.
The expression of phosphor-histone3, an M phase marker, found a significant decrease in the percentage of EdU+pH3+Pax6+ cells among the pH3+Pax6+ cells in cKO mice compared to WT, suggesting a prolonged S to M phase transitions of RGCs.
Further analysis of different cell-cycle phases revealed a specific increase of S-G2-M phase in the absence of Mettl14 but no difference in the G1 phase. Together these results confirm that cKO has a delay in cell-cycle progression and the m6A methylation plays a role in cortical neurogenesis.
In order to confirm that the phenotypes observed were due to m6A and not to Mettl14 depletion, the phenotype of Mettl14 cKO was compared to knockdown of Mettl3. Yoon et al. found a reduction in the percentage of GFP+EdU+NPCs that divided upon Mettl3 KD similar to the effect of Mettl14 cKO. This was further confirmed through EdU fate mapping, which showed similar results between Mettl3 KD and Mettl14. Mettl3 KD and Mettl14 cKO lead to similar phenotypes, demonstrating that decreased levels of m6A results in protracted cell-cycle progression of cortical NPCs and reduced differentiation of RGCs during mouse embryonic cortical neurogenesis.
Yoon et al. (2017) then quantified the levels of nascent mRNA using the 4-thiouridine (4sU) metabolic labeling approach. Lower levels of nascent mRNA of neuronal lineage genes were observed, suggesting that neuronal lineage genes are already expressed in neural stem cells under normal cortical neurogenesis. To confirm this, Tbr2 and Neurod1 protein levels in RGCs were examined and found that Pax6+Neurod1+ cells were more common and widely distributed in cKO mice compared to WT. FT+Pax6+Tbr2+ and FT+Pax6+Neurod1+ cells were also significantly higher in Mettl14 cKO cortex than in WT cortex. Together, these results demonstrate that Mettl14 regulates neuronal lineage gene expression directly in RGCs.
Yoon et al. (2017) also examined whether m6A function is conserved in human cortical neurogenesis, using differentiated human induced pluripotent stem cells (iPSCs) into NESTIN+SOX2+ NPCs. Short hairpin RNA (shRNA) against human METTL14 resulted in a decrease in the percentage of GFP+EdU+ hNPCs that divided with METTL14 KD which suggests delayed cell-cycle progression. The results in humans were similar to those in mice indicating that m6A mRNA methylation plays a conserved role in regulating cortical NPC cell cycle progression in both humans and mice.
This study identified both a critical and conserved role of m6A in the temporal control of mammalian cortical neurogenesis. It plays a part in the promotion of mRNA decay of transcripts related to transcription factors, neural stem cells, the cell cycle and neuronal differentiation. A better understanding of the mechanisms that control neurogenesis could help researchers develop a cure for numerous neurological disorders.
Source: Yoon et al., Temporal Control of Mammalian Cortical Neurogenesis by m6A Methylation. Cell. 171, p 1-13.
