
Cells are continually being generated and eliminated by our bodies in an ongoing cycle crucial for maintaining proper tissue structure and function. The death of cells during normal tissue maintenance results in the release of intracellular DNA material into the bloodstream, where it freely circulates. In fact, healthy individuals can have up to 100 nanograms of non-encapsulated DNA fragments floating around in every milliliter of their blood.1 Circulating cell-free DNA, or cfDNA for short, has proven quite useful in various clinical applications, including the assessment of pregnancy from fetal cfDNA in maternal blood, monitoring of organ transplants from donor cfDNA in the recipient’s blood, and diagnosis of cancer from tumor cfDNA in so-called “liquid biopsies”.
Diseases like diabetes mellitus and multiple sclerosis are characterized by the chronic destruction of particular cell types (β-cells, oligodendrocytes), leading to the loss of critical functions (insulin secretion, MOG glycoprotein production) necessary for life-sustaining biological processes (glucose homeostasis, CNS myelination). Such targeted cell death oftentimes goes undetected in early phases of disease progression when therapeutic intervention would be the most beneficial, before substantial and irreversible loss of tissue function occurs. As these cells die, their genetic contents are expelled into circulation and mix with cfDNA already present from physiological cell turnover.
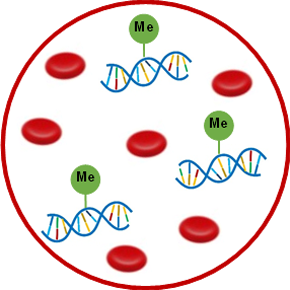
Although all of our body’s cells essentially contain the same genes, the expression of those genes is tissue-specific and tightly regulated by epigenetic factors like DNA methylation and histone modifications. The cfDNA epigenome can thus be utilized to determine the exact source from which the cfDNA originated. For example, hypomethylation of CpG motifs in the insulin promoter distinguishes β-cell-derived cfDNA from non-β-cell DNA flowing through the bloodstream.2 Oligodendrocyte cfDNA in sera from patients with active relapsing-remitting multiple sclerosis is recognizable by its demethylated MOG coding region.3 The ability to uncover and treat pathological conditions before disease onset would certainly improve patient outcomes, and differentially methylated cfDNA can serve as a convenient biomarker for the early detection of disease-associated cell death in vivo.
Processing and analyzing differentially methylated cfDNA is a multistep procedure that generally involves DNA retrieval, chemically modifying the genetic region of interest to discern between methylated and unmethylated sites, and amplifying both the modified and unmodified target sequence to measurable levels. The first step is to efficiently isolate and purify the cfDNA. Current methods for cfDNA isolation are based on the capture of DNA by silicone column binding, phenol-chloroform separation, or silica beads, and tend to be time-consuming and low throughput. Innovative magnetic bead-based size fractionation technology offers a simpler and faster way to obtain high recovery of pure cfDNA. Methylated DNA fragments in particular can be selected for and enriched through methylated DNA immunoprecipitation-based systems employing high quality and non-cross-reactive antibodies.
The next step is to prepare the isolated cfDNA for gene-specific methylation analysis. This commonly entails performing bisulfite conversion, a chemical reaction where unmethylated cytosines are deaminated to uracil while leaving 5-mC intact (DNA methylation covalently adds methyl groups at the 5-carbon position of cytosine rings to form 5-methylcytosine, or 5-mC). As a consequence, methylated and unmethylated cfDNA cytosine residues are differentiated from one another. The traditional conversion method requires a lengthy protocol time (12-16 hours), causing heavy DNA degradation (>80%), high inappropriate 5-mC deamination (>3.5%), and low cytosine conversion rate (<95%).
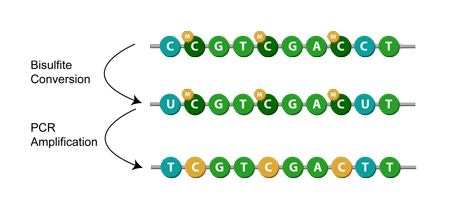
Efforts to condense the entire bisulfite process to just 0.5-1.5 hours have significantly improved cytosine conversion efficiency (>99.9%) and effectively prevent bisulfite-treated DNA degradation. Hydroxymethylated 5-mC, or 5-hmC, is another epigenetic mark of cfDNA garnering research interest4, and recent technology permits the simultaneous identification of both methylated cytosine forms. By coupling bisulfite, directed at unmethylated C residues, with subsequent APOBEC deaminase treatment, which selectively changes 5-mC to thymine, 5-hmC is accordingly discriminated. The converted cfDNA is now ready for more in-depth investigation, typically by PCR and next-generation sequencing methods, to resolve the precise methylation sites within the target region at the single-base resolution level.
Medical procedures can be rather expensive and extremely intrusive, much to the inconvenience and discomfort of the patient. Invasive lumbar punctures to extract cerebrospinal fluid and costly MRIs are physically and financially burdensome to those afflicted with multiple sclerosis, for instance. Differentially methylated cfDNA provides a noninvasive and low-cost alternative to conventional diagnostics. From just a small blood sample, trace amounts of DNA are able to be isolated and identified by their unique epigenome with high degrees of sensitivity and specificity. Methylation-based techniques can be used to pinpoint the origin of cfDNA and assess their blood levels amid the sea of circulating nucleic acid fragments, helping to accurately and precisely diagnose and monitor the relevant pathology. Combined with an epigenetic analytical approach, differentially methylated cfDNA presents new opportunities for clinically approved molecular biomarkers of cell death.
References:
- Esposito A et al. The Emerging Role of “Liquid Biopsies,” Circulating Tumor Cells, and Circulating Cell-Free Tumor DNA in Lung Cancer Diagnosis and Identification of Resistance Mutations. Curr Oncol Rep. 2017, 19:1.
- Akirav EM et a. Detection of β cell death in diabetes using differentially methylated circulating DNA. Proc Natl Acad Sci. 2011, 108:19018-19023.
- Olsen JA et al. A Minimally-invasive Blood-derived Biomarker of Oligodendrocyte Cell-loss in Multiple Sclerosis. EBioMedicine. 2016, 10:227-235.
- Chiu BC et al.Prognostic implications of 5-hydroxymethylcytosines from circulating cell-free DNA in diffuse large B-cell lymphoma. Blood Adv. 2019, 3:2790-2799.
