
The recent COVID-19 pandemic is caused by SARS-CoV-2 (Severe Acute Respiratory Syndrome Coronavirus 2), a novel betacoronavirus. It was first recognized in the beginning of 2020 as a new member of the same coronavirus family that caused SARS and MERS. So far there is no vaccine or effective drugs that can be used for prevention and treatment of COVID-19.
What can we do in this current situation? Other than social distancing and frequent washing of hands, a dietary supplement of the right foods may be an additional approach that can potentially lessen or control the risk of a SARS-Cov-2 infection and help raise your defenses against COVID-19.
SARS-CoV-2 Biology
Before we look at compounds in foods that might be helpful, we first need to understand some of the behind-the-scene mechanics of the virus.
Virus Type: The genome of SARS‐CoV-2 is a positive-sense single‐stranded ribonucleic acid (ssRNA) of approximatly 29700 nucleotides in length, of about 80% identical to that of SARS-CoV-2 and approximately 96% identical to the bat coronavirus BatCoV RaTG13.[1]
Structure of SARS-CoV-2: It has four structural proteins, known as S (spike), E (envelope), M (membrane), and N (nucleocapsid) proteins; the N protein holds the RNA genome, and the S, E, and M proteins together create the viral envelope. The spike protein is the protein responsible for allowing the virus to bind to a host cell through the ACE receptor.
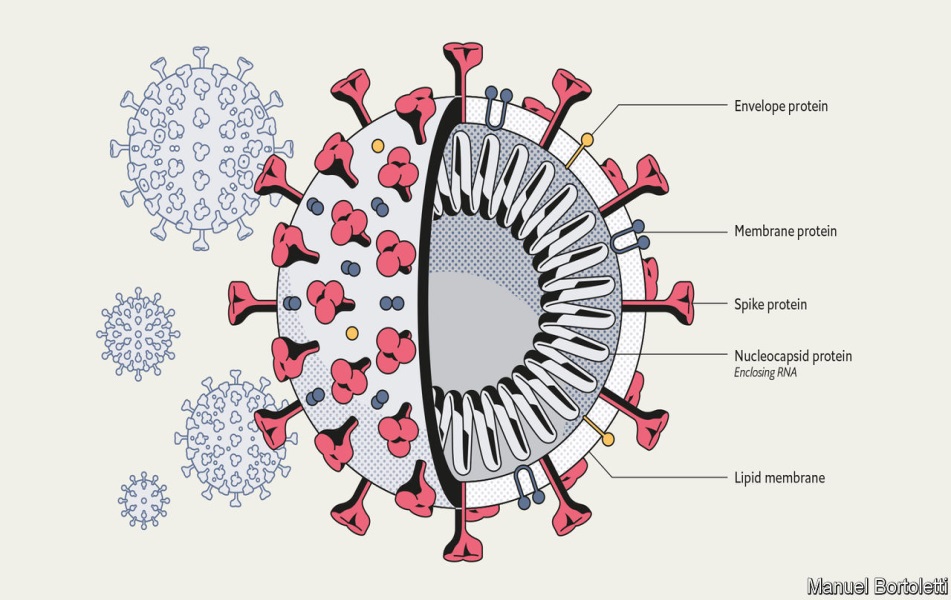
Anatomy of a Killer. Credit: https://www.economist.com/briefing/2020/03/12/understanding-sars-cov-2-and-the-drugs-that-might-lessen-its-power
Cellular Infection Approach: (a) Binding and Fusion: In the case of SARS-CoV-2, the spike glycoprotein (S protein) on the virion surface mediates receptor recognition and membrane fusion. During viral infection, the trimeric S protein is cleaved into S1 and S2 subunits and S1 subunits are released during the transition to the post-fusion conformation.
S1 contains the receptor binding domain (RBD), which directly binds to the peptidase domain (PD) of ACE2[2], while S2 is responsible for membrane fusion. When S1 binds to the host receptor ACE2, another cleavage site on S2 is exposed and is cleaved by host proteases, a process that is critical for viral infection; (b) Replication: SARS-Cov-2 positive ssRNA can function both as a genome and a messenger RNA.
Upon infection, viral RNA synthesis follows the translation and assembly of the viral replicase complexes for replication machinery; (c) Exit from Cells: Following assembly, virions are transported to the cell surface in vesicles and released by exocytosis. The SARS-CoV-2 S protein contains a protease cleavage site and is likely processed by these intracellular proteases during exit from the cell.[3] The viral particles may be therefore ready for entry into the next cell.
Next, we look at how epigenetics plays a role in association with the SARS-CoV-2 virus.
Epigenetic Regulation During SARS-Cov-2 Infection
Epigenetics is the study of heritable changes in gene expression (active versus inactive genes) that do not involve changes to the underlying genome sequence — a change in phenotype without a change in genotype in response to environment stimuli.[4] DNA and RNA methylation are major biological processes in epigenetics.
Epigenetic Regulation of the Virus: Genome modification of RNA viruses by methylation plays an important role in virus replication, assembly, and maturity. For example, the 5′ cap of viral RNA is produced by methylation of the N7-guanosine and 2′-O-adenosine positions. The presence of N6-methyladenosine (m6A) has been described in mRNAs of several ssRNA viruses.[5] In addition, several enzyme/proteins, made by the host and exploited by the virus in the cell membrane fusion and replication could be activated or inactivated by DNA methylation changes.
Epigenetic Modulation of Host Response: Like other ssRNA viruses, when SARS-Cov-2 infects host cells, the virus requires continuous cellular transcription for viral mRNA synthesis. This mechanism implies functional association with the host’s genome expression so that it adapts the infected cell for the host-to-pathogen confrontation at each replication.[6] A key role is that RNA viruses can recruit host DNA methytransferases (DNMTs) to methylate and decrease gene function of specific genes including those for shaping innate and adaptive immune responses.
Targeted Sites for Cellular Virus Infection Inhibition
The following enzymes/proteins from either the host or the virus could be targets for inhibiting or reducing viral infection.
1. ACE-2 Receptor.Responsible for SARS-COV-2 binding to cells.
2. Spike Protein. Mediates virus binding and cell entry through receptor-binding domain (RBD).
3. Protease Furin and TMPRSS2. It was found that S protein of SARS-COV-2 contains the furin cleavage site (PRRARSV) and TMPRSS2 cleavage site (S1/S2 R682/683/685 and S2 R815), which are responsible for S protein priming and cell membrane fusion. Specifically, the furin site in SARS-Cov-2 S1/S2 boundary is unique and critical for highly efficient cleavage of the S protein and is an important determinant of the transmissibility and pathogenicity of the virus.
4. Replicase Complexes. Includes protease 3CLpro, PLpro, helicase, and RNA dependent RNA polymerase (RdRP), which are responsible for virus replication after entry into cells.
5. DNA Methyltransferases (DNMTs). Responsible for DNA methylation to suppress gene function. Inhibition of DNMTs will re-activate immunity-related gene function responsible for combatting viral infection.
6. RNA Methyltransferases. Includes enzymes such as METTL3/14 that may facilitate RdRP stability and mRNA cap guanine-N7-methyltransferase that is responsible for protecting the virus’s ability to replicate.
7. Proteins for Virus Escaping from Innate Immunity. May include Nsp1, NsP3c, and ORF7a.
Epigenetic Prevention/Control by Epigenetic Diet
Oranges, onions, berries, and tea —
Epigenetic diet for me.
Having them plenty every day,
COVID-19 is kept away.
Certain foods, especially fruits that contain abundant bioactive compounds, have been shown to modify the epigenome leading to beneficial health outcomes including anti-viral effects. Specifically, to help reduce risk of infection by SARS-Cov-2, one should take note of the following: (a) the fruits should contain the most effective compounds against the virus through proven enzyme/protein targets; and (b) sufficient amount of such compounds should be obtained from the diet in order to reach an effective concentration in the body to suppress viral infection. Based on this principle and scientific literature, several flavonoids have been selected and such compounds could be potentially the most helpful in reducing infection risk:
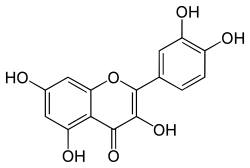
Quercetin (depicted above) and its derivatives are flavonols and most abundant in onion and juniper berries. The target enzyme/proteins by quercetin are ACE-2, 3CLpro, and helicase. Molecular docking indicates a good interaction between quercetin and these proteins. It was reported that quercetin is quite effective against the SARS virus.[7]

Compounds Naringenin, Luteolin, Hesperetin, Hesperidin (depicted above in respective order) — are flavonones mainly found in citrus fruits and are the most abundant in oranges. Luteolin is also quite rich in juniper berries. Molecular docking shows these compounds can interact well with ACE-2, furin, 3CLpro, and/or the virus S protein.[8]
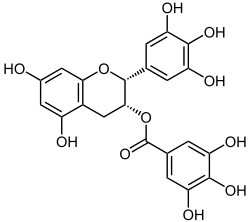
EGCG (epigallocatechin-3-gallate) is a specific flavanol derivative, flavan-3-ol. Green tea contains the most EGCG. Molecular docking shows EGCG can interact well with DNMTs, ACE-2, and helicase.[9]
Listed in Table 1 are key compound contents in the aforementioned foods, as well as target enzymes/proteins, EC50 (effective concentration to inhibit 50% of viral or key protein activity), PPC (peak plasma concentration), and consumption amount needed to achieve the PPC after intake of these compounds.[10-16]
Table 1:
| Epigenetic Compound | Target Site | Content (mg/100 g) | EC50* (uM) | PPC (uM) | Amount to Achieve PPC (mg) |
| Onion | |||||
| Quercetin (derivative) | ACE2 3CLpro Helicase | 51.8 (6330) | 8.1 | 11.8 | 150 |
| Oranges | |||||
| Luteolin | ACE2 Furin | 0.19 | 3.6 | 0.43 | 150 |
| Hesperetin | ACE2 3CLpro | 21.8 | 8.3 | 2.7 | 135 |
| Hesperidin | ACE2 3CLpro | 40 | 8 | 2.2 | 126 |
| Naringenin | ACE2 3CLpro | 4 | 52 | 7.1 | 135 |
| Green Tea | |||||
| EGCG** | DNMT S protein-ACE2 binding Helicase | 7384 |
0.4 4.3 0.3 | 0.71 | 73 |
| Juniper Berries (ripe) | |||||
| Quercetin | ACE2 3CLpro Helicase | 46.6 | 8.1 | 11.8 | 150 |
| Luteolin | ACE2 Furin | 69 | 3.6 | 0.6 | 35.2 |
Table 1. List of potentially effective foods and their key compound(s) that may reduce risk of SARS-CoV-2 infection; * = Coronavirus or critical protein inhibition; ** = 0.4 is for DNMTs, 4.3 for coronavirus inhibition, and 0.3 for viral helicase.
Amount of Food to Consume
The inhibition of viral infection is dependent on the plasma concentration of the inhibitors. In general, 300 g of fresh red onion, 300 ml of squeezed orange juice, 1 g of green tea, and 50 g juniper berries would achieve the necessary PPC of quercetin, hesperidin, EGCG, and luteolin, respectively as described in Table 1.
Quercetin alone could achieve a PPC for 50% virus inhibition with 150 mg amount intake, which could be obtained from 300 g of onion. EGCG alone may also have a good inhibitory effect on the virus with a sufficient amount (i.e., 400 mg), as low EC50 for the virus can be achieved. Other compounds alone may not be sufficient to significantly inhibit the virus. However, an additive effect to strongly suppress the virus could be achieved with these compounds together as they target the same enzyme/proteins.
The active compounds from orange and juniper berries can be simply obtained by eating these fruits directly, or drinking the juice from them. The best cooking method for onions is light frying or sautéing that will retain nearly 100% of quercetin for intake. To maximize EGCG from green tea, the best method would be to using 5 g of green tea in 300 ml of water, brewed at 85 degrees Celsius for 3 minutes. EGCG content is expected to reach a maximum of 50 mg per 100 ml of water.
Eating the right foods is an easy and convenient way strengthen your defenses against this virus with a diet that is readily available.
Disclaimer: This article is not intended for clinical or diagnostic applications and its information is implied through scientific evidence, and may contain points that represent a speculative opinion of the author based on related scholarly publications on in vitro animal and human studies.
Selected References:
- Paraskevis D et al. (2020) Full-genome evolutionary analysis of the novel corona virus (2019-nCoV) rejects the hypothesis of emergence as a result of a recent recombination event. Infect Genet Evol. 79.
- Walls AC et al. (2020) Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein. Cell.
- Coutard B et al. (2020) The spike glycoprotein of the new coronavirus 2019-nCoV contains a furin-like cleavage site absent in CoV of the same clade. Antiviral Res. 176.
- WhatIsEpigenetics. Epigenetics: Fundamentals, History, and Examples. Web.
- Wu L et al. (2019) Virology Journal. 16.
- Marcos-Villar L et al. (2018) Epigenetic control of influenza virus: role of H3K79 methylation in interferon-induced antiviral response Scientific Reports. 8.
- World Health Organization. Hospital infection control guidance for severe acute respiratory syndrome (SARS) 24 April, 2003. Web.
- Utomo RY et al. (2020) Revealing the Potency of Citrus and Galangal Constituents to Halt SARS-CoV-2 Infection. Preprints.
- Jo S et al. (2020) J of Enz Inhibition and Med Chem. 1.
- Kwak JH et al. (2017) Saudi J of Biol Sci. 24 (6).
- Kumar D et al. (2019) Mechanistic insights into Zika virus NS3 helicase inhibition by Epigallocatechin-3-gallate. bioRxiv.
- McAnlis GT et al. (1999) Absorption and antioxidant effects of quercetin from onions, in man. Eur J of Clinic Nutrition. 53 (2):92-6.
- Palmer PA. (2018) https://www.fda.gov/media/133322/download.
- Yi, L et al. (2004) Small Molecules Blocking the Entry of Severe Acute Respiratory Syndrome Coronavirus into Host. Cells J Virol. 78 (20).
- Guerrero L et al. (2012) Inhibition of Angiotensin-Converting Enzyme Activity by Flavonoids: Structure-Activity Relationship Studies. PLoS One. 7 (11).
- Matsumoto M et al. (2005) Inhibitory effects of epigallocatechin gallate on the propagation of bovine coronavirus in Madin-Darby bovine kidney cells. Animal Science Journal. 76 (5):507 – 512.
