DNA bisulfite conversion is a unique tool used to discriminate between methylated and unmethylated cytosines for DNA methylation studies. Only bisulfite modification of DNA followed by sequencing yields reliable information on the methylation states of individual cytosines at single base resolution. The bisulfite modification technique uses bisulfite salt to deaminate cytosine residues on single-stranded DNA, converting them to uracil while leaving 5-methylcytosine unchanged (Figure 1).
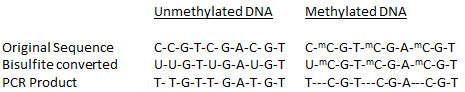
To effectively and efficiently prepare converted DNA for use in various downstream analyses, an ideal DNA bisulfite modification method should be: (1) highly accurate to allow for the complete conversion of cytosine to uracil (correct conversion) without deamination of methylcytosine to thymine (incorrect conversion); (2) conservative enough to allow for minimal DNA degradation and (3) rapid enough to enable the bisulfite process to be as short as possible.
Here we have highlighted some of the key considerations for successful bisulfite conversion based on feedback from scientists at Epigentek, who have extensive experience and expertise with the modification process. Some common pitfalls can be easily avoided to not only improve your results in downstream applications but to save time, precious samples and your sanity.
Bisulfite Conversion – Tips and Techniques
Before performing the bisulfite conversion reaction, consider your downstream application. Different methods result in different post-conversion fragment sizes which may or may not be suitable for your next step. High heat combined with a short incubation time can save a lot of time in the conversion process but may also result in shorter fragment sizes and more DNA degradation than lower heat/variable cycling temperatures and a longer incubation time (Table 1).
As mentioned above, the bisulfite process itself will fragment your DNA due to acidic conditions, reaction time and temperature. Therefore, the quality of the input DNA should be relatively high. Check that your input DNA is not degraded by running a gel. The DNA 260/280 ratio should be 1.6-1.9. For optimal modification and sufficient DNA for downstream applications, use 50-200 ng for conversion.
Proper storage and shelf-life of reagents may seem obvious but let’s admit it – we’ve all tried to push the limits using questionable reagents in hopes of success. For optimal performance, prepare fresh modification solution or ensure that previously prepared DNA modification solution has been stored at ‑20˚C away from light, for no more than 2 weeks.
Poorly modified DNA may be the result of template DNA containing high GC-rich regions or secondary structures. In such cases the bisulfite reaction time may be extended. However, make sure that the reaction temperature it not too high which could result in excessive DNA degradation and considerably low yields after purification.
Desulphonation, the final step in the conversion process, is critical for successful downstream PCR applications. Insufficient desulphonation may result in incomplete conversion of the uracil-sulphonate reaction intermediates to uracil, thus inhibiting DNA polymerase from replicating template DNA. To address this issue, use freshly prepared ethanol solutions to make desulphonation solutions and ensure that the desulphonation step is carried out to completion.
Post-bisulfite QC
A quick evaluation can give you information about the quality and relative quantity of your bisulfite converted DNA. The advantage of using these techniques is that they use very little (1-3 µl) of your bisulfite converted DNA, reserving the majority of your sample for your next application.
For qualitative analysis, a real-time PCR assay using primers for unmethylated regions such as beta-actin or GAPDH or that avoid CpG sites can tell you if the DNA quality is suitable or not for PCR.
For quantitative analysis, the amount of bisulfite DNA may be measured using a fluorescence detection method such as that used for RNA – since the DNA is now single-stranded and the majority of cytosines have been converted to uracil. Measure your converted DNA relative to a blank or RNA standard to estimate your yield.
Below is a table highlighting some of Epigentek’s most popular bisulfite modification kits. The varied selection makes it easy to choose a kit to meet any specific research requirement.
Table 1. Bisulfite Conversion Kits for Every Research Need and Commercial Platform
|
Methylamp DNA Modification Kit |
Methylamp Whole Cell Bisulfite Modification Kit |
BisulFlash DNA Modification Kit |
BisulFlash DNA Bisulfite Conversion Mag-96 Kit |
|
| Catalog Number |
P-1001 |
P-1016 |
P-1026 |
P-1050 |
| Purpose |
Basic bisulfite conversion; |
Directly bisulfite convert DNA in cells or tissue – no need for pre-isolation of your DNA |
Rapid DNA bisulfite conversion; denature and modify in one 30 minute step |
Fast bisulfite conversion using a liquid bisulfite reagent and magnetic bead cleanup. High-throughput format |
Properties
| Starting Material |
DNA |
cells, tissues, blood |
DNA |
DNA |
| Lowest Amount of Starting Material |
50 pg |
≤ 100 cells/≤ 1 ul blood |
200 pg |
0.1 µg |
| Bisulfite Processing Time |
1.5 h |
1.5 h |
20 mins |
45 mins |
| Total Processing Time |
≤ 2 h |
< 3 h |
1.5 h |
< 1.5 h |
| Post-conversion DNA Fragment Size |
200-2000 bp; Avg 800 bp |
200-2000 bp; Avg 800 bp |
100-400 bp; Avg 250 bp |
200-2000 bp; Avg 800 bp |
| Conversion Efficiency |
99.90% |
99.90% |
> 99.9% |
> 99.9% |
| Bisulfite DNA Recovery |
>75% |
>85% |
75% |
>75% |
Downstream Application
| Methylation Microarray (Illumina Infinium) |
✓ |
✓ |
✓ |
✓ |
| Illumina NGS |
✓ |
✓ |
✓ |
✓ |
| Conventional MS-PCR |
✓ |
✓ |
✓ |
|
| Real-time PCR MS-PCR |
✓ |
✓ |
✓ |
✓ |
| Pyrosequencing |
✓ |
✓ |
✓ |
✓ |
