A non-coding RNA (ncRNA) is a functional RNA molecule that is transcribed from DNA but not translated into proteins. Epigenetic related ncRNAs include miRNA, siRNA, piRNA and lncRNA. In general, ncRNAs function to regulate gene expression at the transcriptional and post-transcriptional level. Those ncRNAs that appear to be involved in epigenetic processes can be divided into two main groups; the short ncRNAs (<30 nts) and the long ncRNAs (>200 nts). The three major classes of short non-coding RNAs are microRNAs (miRNAs), short interfering RNAs (siRNAs), and piwi-interacting RNAs (piRNAs). Both major groups are shown to play a role in heterochromatin formation, histone modification, DNA methylation targeting, and gene silencing.
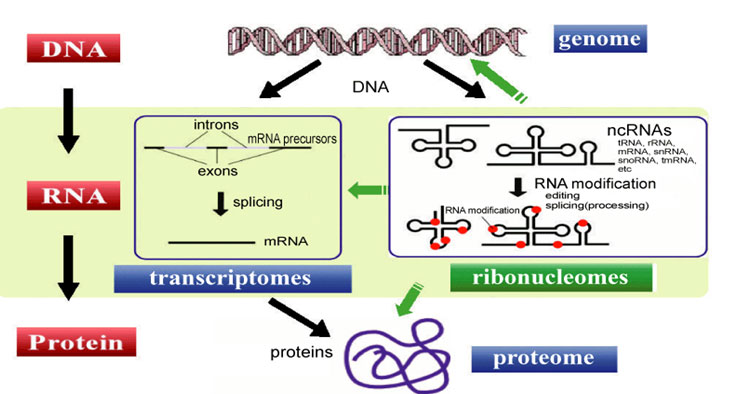
Short ncRNAs
MicroRNAs (miRNA) generally bind to a specific target messenger RNA with a complementary sequence to induce cleavage, or degradation or block translation. This may be done in the context of a feedback mechanism that involves chromosome methylation. For example, miRNA genes mir-127 and mir-136 were found to be involved in regulating the genetic imprinting of Rtl1, a key gene involved in placenta formation in mice. Methylation of a specific region in the paternal chromosome results in expression of Rtl1. If the chromosome is not methylated, as on the maternal chromosome, mir-127 and mir-136 are produced and bind to the Rtl1 transcript and induce degradation. Lack of Rtl1 protein expression due to improper epigenetic modifications can result in fetal death in mice.12
Short interfering RNAs (siRNA) function in a similar way as miRNAs to mediate post-transcriptional gene silencing (PTGS) as a result of mRNA degradation. In addition to this function, siRNAs have also been shown to induce heterochromatin formation via an RNA-induced transcriptional silencing (RITS) complex which when bound to siRNA promotes H3K9 methylation and chromatin condensation.3
Piwi-interacting RNAs (piRNA) are so named due to their interaction with the piwi family of proteins. The primary function of these RNA molecules involves chromatin regulation and suppression of transposon activity in germline and somatic cells. PiRNAs that are antisense to expressed transposons target and cleave the transposon in complexes with PIWI-proteins. This cleavage generates additional piRNAs which target and cleave additional transposons. This cycle continues to produce an abundance of piRNAs and augment transposon silencing.45
Long ncRNAs
Many lncRNAs can complex with chromatin-modifying proteins and recruit their catalytic activity to specific sites in the genome, thereby modifying chromatin states and influencing gene expression.6 The majority of non-coding RNA transcripts belong to the group lncRNAs. Long ncRNAs function in chromatin remodeling, transcriptional regulation, post-transcriptional regulation, and as precursors for siRNAs.7 One particular subgroup of lncRNAs, the large intergenic non-coding RNAs (lincRNAs), has been associated with chromatin modifying complexes which can target specific genomic loci to promote specific epigenetic states.8 One widely known example of this is the role of X-inactive specific transcript gene (Xist), in X-chromosome inactivation (XCI). This process involves two lncRNAs; Xist and its antisense transcript Tsix, a negative regulator of Xist. Prior to differentiation, Xist and Tsix are actively transcribed due to H3K4 dimethylation of the Xist gene. In this state XCI is a random event. Upon differentiation, Xist expression is elevated resulting in Xist RNA coating the future inactive X chromosome which triggers extensive histone methylation and chromosome inactivation.9
Need a refresher on epigenetics again? Read on: Fundamentals of Epigenetics
