Bisulfite sequencing offers researchers a profound look into the epigenome and the methylation status of genes. A significant driving force in the development of epigenetic research since 1992, the detection of CpG methylation and methylation abnormalities in DNA via bisulfite sequencing has become overwhelmingly popular – and interest continues to grow.
The unparalleled power of next-generation sequencing (NGS) platforms provides researchers with new insights into the nuances of gene expression and countless other critical cellular processes. Still, bioinformatics expertise, cost, and throughput limitations prevent NGS bisulfite sequencing research from achieving ultimate accessibility and utilization by a range of scientists.
Developing technologies, however, such as single molecule real-time (SMRT) DNA sequencing platforms and other progressive tools, including commercially available bisulfite conversion kits and the novel SMRT bisulfite sequencing (SMRT-BS) method, are paving the way for researchers to quantitatively sequence longer read lengths, at a lower cost, and at high-throughput capacities. In light of these progressions, the practicality and accuracy of targeted bisulfite sequencing using SMRT-BS could take epigenetic research in a once unimaginable direction.
Bisulfite Sequencing Methods
Next-generation bisulfite sequencing enables researchers to profile DNA methylation across an entire genome at an unmatched resolution. But next-generation bisulfite sequencing approaches such as reduced representation bisulfite sequencing (RRBS) and whole-genome bisulfite sequencing, although popularly used to detect CpG methylation in a single experiment, are limited.
These approaches are expensive per sample, restricted to low throughput, and require in-depth bioinformatics knowledge, therefore prompting many researchers to actually continue to utilize earlier targeted bisulfite sequencing techniques, which allows a more focused interrogation of a region of interest. Unfortunately, the limitations of targeted bisulfite seq are still constricting and offer little room for long reads and multiplexing. But what if these restrictions could be overcome?
A recent development in bisulfite conversion methods, known as single molecule real-time bisulfite sequencing (SMRT-BS) could do just that.
This newly developed protocol, along with commercially available kits and SMRT sequencing platforms such as the Pacific Bio system, could potentially elevate the abilities of both large and small-scale labs to conduct cost-effective, extraordinarily accurate, quantitative CpG methylation detection analysis on a scale that many have thought to be previously unattainable.
SMRT-BS
The single molecule real-time bisulfite sequencing (SMRT-BS) method was presented at ASHG in 2014 by Stuart A. Scott, PhD, from the Department of Genetics and Genomic Sciences at the Icahn School of Medicine at Mount Sinai. In an attempt to overcome the short read length and multiplexing limitations of targeted bisulfite sequencing, a team of researchers developed an innovative technique that combines bisulfite conversion with third-generation single molecule real-time (SMRT) sequencing without the need for library preparation.
A significant advantage of this SMRT-BS method comes from its powerful insight into DNA methylation analysis using limited sample input. SMRT sequencing technology in general is capable of directly detecting base modifications without the need for bisulfite conversion. However, because amplification of DNA would erase base modifications, the SMRT sequencing method requires relatively large amounts of native, unamplified DNA as input material. For many, an abundance of sample material is often difficult to come by. But researchers can overcome this limitation with the proposed SMRT-BS method, which allows for amplification of small amounts of DNA and still takes advantage of PacBio’s long-range read capabilities. Amplifying low quantities of just 50-100 ng of DNA as opposed to inserting 5-10 µg of native DNA and still obtaining accurate results is a significant advantage and much more practical, especially for those who are restricted by limited sample amounts.
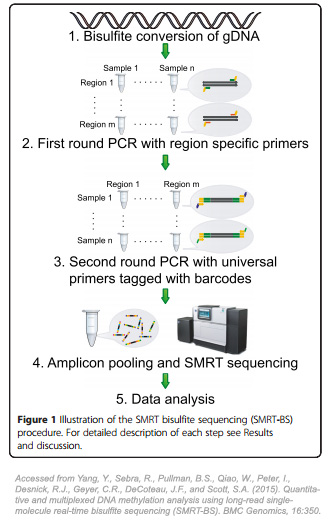
In the SMRT-BS procedure described in the study published in BMC Genomics, genomic DNA is first subjected to bisulfite conversion. Then, primers that are region-specific along with universal oligonucleotide tags are used to amplify the bisulfite-treated DNA. Universal anti-tag primers in addition to sample-specific multiplexing barcodes then re-amplify amplicon templates. After amplicon purification, pooling, and SMRT sequencing with the PacBio RS II system, CpG methylation can then be quantitated (Fig. 1).
After testing six commercially available bisulfite conversion kits in the study, the researchers found that only one, Epigentek’s Methylamp DNA Modification Kit, was most ideal, producing consistent amplification of the longer 1109 bp product at 65°C and amplifying products up to an impressive ~2.0 kb.
Typically, high throughput methods face limitations of amplicon size and short read length due to the harsh chemical fragmentation of DNA during bisulfite conversion, but the most robust amplification of the longer 1109 bp amplicon could be achieved using bisulfite-converted DNA from the Methylamp kit. Researchers also ensured that this was not due to poor bisulfite conversion efficiency.
This optimal, long-range amplification protocol allowed the team to attain significant amplification of DNA products up to ~2.0 kb, with the most consistently and stably amplified product at ~1.5 kb. Because PacBio’s system is capable of sequencing reads over 40,000 base pairs in comparison to Illumina sequencing technology which, at most, sequences amplicons 300 bp in length, it is highly suited to this developing bisulfite sequencing application alongside the Methylamp kit with its capacity to produce intact long fragment lengths.
SMRT-BS Validation and Reproducibility
Multiple analyses validate and strongly support the impressive accomplishments of this cost-effective targeted bisulfite sequencing approach, accomplishments which include the ability to sequence amplicons ~1.5 kb in length and, theoretically, ~91% of all CpG islands in the human genome.
The researchers compared methylation levels from peripheral blood DNA of TUBGCP3, MEST, and EHPA8 CpG islands with two unique and independent second-generation sequencing and microarray platforms and found that the CpG methylation levels were correlated impressively with the SMRT-BS technology, r = 0.906 ± 0.052 and r = 0.933 ± 0.031, respectively. For all methylation levels and amplicon sizes (625-1491 bp), reproducibility was very high. Between independent triplicate amplicons, the researchers found an average overall correlation of r = 0.972 ± 0.024.
They also subjected four CpG island amplicons and 30 hematological malignancy cell lines to the new method and concluded that SMRT-BS is also highly accurate and comparable to current methods. For instance, one of the cell lines subjected to SMRT-BS with a sequencing depth of 101X had also been investigated using RRBS methylation analysis with a sequencing depth of 31X through the ENCODE project. Among 129 CpG sites, 78 CpG sites had >20X coverage by RRBS and the Pearson correlation between the methylation levels of these two methods at these CpG sites was an exceptional 0.900.
SMRT-BS Considerations
Although this exciting methodology may open new doors for the expansion of epigenetic research publications for a wide range of researchers, there are some considerations that should be acknowledged.
Scientists should note that reduced sequencing depth was associated with increased variability in CpG methylation quantitation. This was especially applicable to CpG regions with intermediate methylation. Therefore, researchers should tactfully choose sequencing depth in order to achieve optimal results with SMRT-BS. Overall, reduced correlation was observed with amplicons greater than 1.0 kb compared to the correlation of amplicons less than 1.0 kb and because greater sequencing depth was linked to greater methylation reproducibility and shorter amplicon length, the researchers suggest overcoming the increased variability by increasing sequencing depth for regions with intermediate methylation levels. This adjustment is expected to achieve similar confidence intervals and margins of error for large amplicons.
Additionally, researchers should be selective with their bisulfite conversion kit of choice for SMRT-BS, as only the most optimal bisulfite PCR protocol was successfully utilized in this study and very few kits tested produced desirable bisulfite conversion rates or amplicon lengths. The use of a third-generation sequencing platform such as the PacBio RS II system allows for extraordinarily long reads and accurate sequencing of DNA, but the quality and appropriate length of the DNA fragments must be initially achieved for successful results.
Future Directions with SMRT-BS
With the exciting development of SMRT-BS and long-read multiplexed amplicon sequencing, researchers can interrogate the genome at an extraordinary depth and perform a closer examination of a specific region of interest at a lower cost. Combining a high quality bisulfite conversion kit with the exceptional sequencing power of the SMRT-based sequencing platform, such as the PacBio RS II system, will enable SMRT-BS to raise epigenetic research to new heights as a highly accurate, cost-effective, multiplexed bisulfite sequencing method for targeted CpG methylation analysis.
Reference:
Yang, Y., Sebra, R., Pullman, B.S., Qiao, W., Peter, I., Desnick, R.J., Geyer, C.R., DeCoteau, J.F., and Scott, S.A. (2015). Quantitative and multiplexed DNA methylation analysis using long-read single-molecule real-time bisulfite sequencing (SMRT-BS). BMC Genomics, 16:350.