Every epigenetics scientist knows that chromatin immunoprecipitation (ChIP) is a valuable technique for studying protein-DNA interactions. They also know that antibodies used in ChIP to capture the DNA/protein complex must be reliable and specifically recognize the fixed protein that is bound to the chromatin complex. But how can a researcher be certain that the antibodies they are using work well in ChIP? For many researchers it is not always the case to be “blessed” with a quality ChIP-grade antibody and, unfortunately, this may be the defining difference for a successful ChIP experiment.
We’ve already covered some quick tips to validate your antibody for ChIP experiments. But, the typical course of action – performing labor-intensive and time-consuming ChIP protocol to test the antibody – takes just as long as the actual ChIP experiment and you have no choice but to carry out full-blown PCR amplification and sequencing. Thankfully, the scientists at Epigentek recognized this challenge and recently developed their easy-to-use Pre-Sure™ ChIP Antibody Validation Kit. It lets you quickly validate your antibody for use in ChIP – so you don’t waste time redoing the entire ChIP protocol when your antibody fails.
“Our clients would call with technical questions regarding ChIP and on many occasions their experiment didn’t work out because of the antibody,” says Milka Rodriguez, Ph.D., a scientist at Epigentek who helped develop the kit. “We wanted to offer the researcher a way to validate antibodies in their own lab, so that they can concentrate on their experiment and get high quality, reliable results. Researchers spend enough time in the lab already – this is a must.”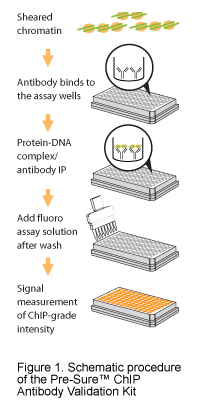
When antibodies are labeled “ChIP-grade” or “ChIP-qualified” from suppliers, they often rely on the assumption that the more complementary applications in which the antibody is successful (Western blot, immunofluorescence, peptide arrays, and peptide competition assays), the more likely the antibody is to work in ChIP. This widely-held assumption, however, doesn’t always ring true. In fact, determining whether an antibody is ChIP-grade completely depends on if, and how well, it captures specific protein-DNA complexes.
William Lee, Vice President of Operations at Epigentek, explains, “The market is saturated with questionable ChIP-grade claims of antibodies due to improper use of Western blot and immunohistochemistry data as the validation basis, both of which are actually unreliable applications for ChIP determination.”
Many antibodies are not validated for detecting enriched target protein/DNA complexes contained in chromatin – a process which requires the antibodies to have high affinity and sensitivity in order to avoid grabbing the “junk.” Careful validation of antibodies for ChIP is important to the integrity and success of the research study. Epigentek’s Pre-Sure kit directly quantifies the protein-DNA complex, making it accurate in determining whether the antibody is ChIP-grade and avoiding the false negative possibilities that may occur with ChIP-PCR validation (no binding at genomic locations investigated or improper primers). Epigentek hopes to place the power to validate antibodies in the hands of the researcher so he or she can attain high quality ChIP results.
Until now, there were no assays or tests available to quickly check if the target protein/DNA complex was successfully enriched by the antibody. This new kit offers an easier, quicker, and accurate way to validate your antibody for ChIP.
It takes only three hours to measure your antibodies’ ChIP-grade intensity and determine whether the antibody enriched the target protein/DNA successfully. Using a fluorescence-based microplate reader, there’s no need for cumbersome PCR amplification and sequencing. Not only that, you can test 43 antibodies at once and get results that are entirely consistent with results you’d get from running ChIP-qPCR and ChIP-seq.
For more info on Epigentek’s ChIP antibody validation kit, visit: http://www.epigentek.com/catalog/pre-sure-chip-antibody-validation-kit-p-3797.html.

